Animal model
Wild-type C57BL/6J mice were purchased from the Model Animal Research Center of Nanjing University. Male mice were exclusively used for all experiments conducted in this study. These mice were housed in a controlled environment with a 12-h light/dark cycle (illuminated from 7:00) and granted unrestricted access to both food and water. All experiments were approved by the Animal Ethical and Welfare Committee of Nanjing University. The approved animal ethical and welfare number is IACUC-2010001. All animal care and handling procedures were performed in accordance with National Institutes of Health Guidelines for the Care and Use of Laboratory Animals.
Plasma and sEV isolation
Human blood was extracted from each donor by aseptic venipuncture and collected into a vacuum tube (~5 ml) containing EDTA as an anticoagulant. Mouse blood was collected by intracardiac puncture into an Eppendorf tube (~1.5 ml) containing trisodium citrate as an anticoagulant. Plasma was obtained by centrifuging whole blood at room temperature for 15 min at 1,600g. The purified plasma samples were immediately centrifuged to obtain the purified sEVs or stored at −80 °C.
Plasma sEVs were isolated by ultracentrifugation. In brief, the plasma was diluted with an equal volume of PBS and subjected to a series of centrifugation steps at different speeds and durations. Specifically, centrifugation was performed at 500g for 5 min at 4 °C, followed by 3,000g for 25 min at 4 °C, 12,000g for 60 min at 4 °C and, finally, 120,000g for 70 min at 4 °C (polycarbonate tubes, 355630, Beckman Coulter; MLA-55 rotor, Optima MAX-XP, Beckman Coulter). The sEV pellet was collected, and the remaining supernatant was further processed using the Total Exosome Isolation Kit (from plasma) (Invitrogen, 4484450) according to the manufacturer’s instructions. In brief, 0.2 volumes of Exosome Precipitation Reagent were added to the supernatant, which was then incubated at room temperature for 10 min. Subsequently, sEVs and the supernatant were separated via centrifugation at 10,000g for 30 min at 4 °C, with the resulting sEV pellets being collected and resuspended in PBS.
The yield of sEVs was assessed by quantifying the total protein content using the Bicinchoninic Acid (BCA) Kit (Thermo Fisher Scientific). Approximately 1,000 μg of total protein content of sEVs could be obtained from 1 ml of plasma. Because the recovery yield of sEVs from viscous body fluids (plasma and serum) using ultracentrifugation-based procedures has been estimated to range from 5% to 25%60, the genuine sEV content in plasma might be several folds higher than the measured value (1.00 μg of total protein per microliter). To simulate the natural concentration of sEVs in plasma, the volume of PBS solution used to dissolve sEVs was approximately 0.56 times the original volume of plasma, so the final concentration of sEVs in PBS was generally 1.80 μg of total protein per microliter. The sEV samples were promptly stored at 4 °C for short-term storage and at −20 °C for long-term storage.
The size and concentration of sEVs were analyzed using a NanoSight LM10 system, as previously described61. The morphology and size of sEVs were also examined using a Hitachi TEM system (Hitachi, HT7700), as previously described61. To analyze the purity and protein contents of sEVs, the sEVs supplemented with RIPA lysis buffer were subjected to western blotting for the classical sEV markers (CD63, ~50 kD; TSG101, ~47 kD; Alix, ~100 kD; and CD9, ~25 kD), along with the major plasma protein albumin (~70 kD) and the endoplasmic reticulum protein calnexin (~55 kD). To analyze the RNA contents of sEVs, total RNA was isolated from sEVs using TRIzol reagent, and the levels of sEV miRNA were quantified.
Injection of sEVs
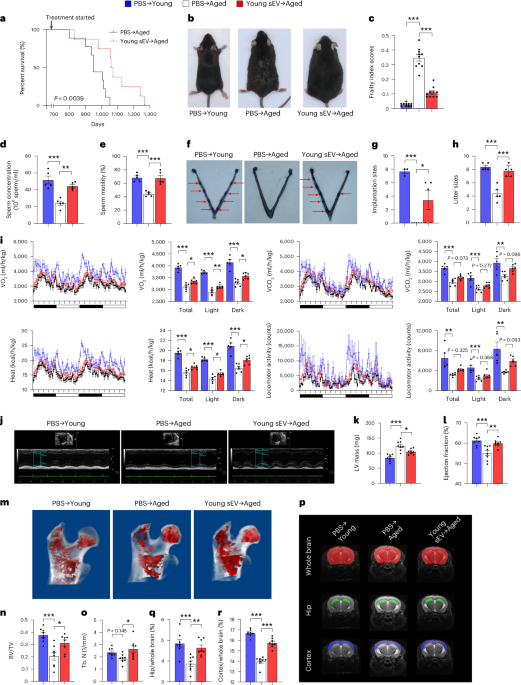
In long-term experiments, young sEVs were isolated from the plasma of 2-month-old male mice using ultracentrifugation and subsequently diluted in PBS to achieve a concentration of 1.80 μg of total protein per microliter. Aged male mice (20 months) were intravenously injected with 200 μl of PBS or young sEVs once a week. At different timepoints, changes of physiological activities and functions, including sperm quality and male fertility (22-month-old mice receiving eight injections); metabolic rate and energy expenditure (23-month-old mice receiving 12 injections); cardiac functions (23.5-month-old mice receiving 14 injections); bone microarchitecture (24-month-old mice receiving 16 injections); and brain volume alterations (24.5-month-old mice receiving 18 injections), were evaluated. As a control, a group of young male mice (2 months) receiving only PBS was simultaneously monitored.
In short-term experiments, sEVs were isolated from the plasma of 2-month-old or 20-month-old male mice using ultracentrifugation. These sEVs were then diluted in PBS to achieve a concentration of 1.80 μg of total protein per microliter. Aged male mice (21 months) were intravenously injected with 200 μl of PBS or young sEVs seven times over 2 weeks, and young male mice (2 months) were intravenously injected with 200 μl of PBS or aged sEVs seven times over 2 weeks. PBS was injected at the same volume (200 μl) and frequency (seven times over 2 weeks) as a control. Moreover, sEVs were purified from the plasma of young male donors (19–24 years) and intravenously injected into aged mice (21 months) at the same dose (1.80 μg of total protein per microliter, 200 μl) and frequency (seven times over 2 weeks) as mouse sEVs. Subsequently, these mice underwent a battery of behavioral tests to evaluate memory and endurance performance, or they were evaluated using multiple senescence biomarkers to assess senescent phenotypes.
Lifespan
The aging mouse cohorts were accommodated in the animal core facility at Nanjing University and granted unrestricted access to a standard laboratory diet and water. Mice allocated for the survival study were exclusively designated for this purpose and were not subjected to any other biochemical, physiological or metabolic analyses. Aged mice received intravenous injections of an equal volume (200 μl) of either PBS or young sEVs once a week. These mice were carefully monitored daily until their natural death.
Fertility assessment
Aged males injected with PBS or young sEVs were mated with 4-month-old female mice. Female mice were proven to be fertile after a minimum of one litter. To conduct a continuous mating study, male and female pairs were co-housed for a 30-d period. Each morning, the females were examined for the presence of copulation plugs, pregnancy and litter size. The presence of a vaginal plug was considered indicative of successful mating within the pairs. The number of offspring per litter was documented using a camera (Canon). To visualize implantation sites in the uterus, males and females were paired overnight. The presence of a vaginal plug the next morning signified successful mating, and the midpoint of that day was designated as 0.5 dpc. The embryo implantation was examined at 4.5 dpc in pregnant mice. Approximately 5 min after intravenously injecting 200 µl of 1% Evans Blue per mouse, the mice were euthanized, and pictures of the incised uterus were taken. The blue bands were recorded and counted as the implantation site.
Quantification of testosterone levels by ELISA
Testosterone levels were quantified using ELISA kits (Thermo Fisher Scientific) following the manufacturer’s instructions. In brief, mouse plasma samples were collected in 1.5-ml centrifuge tubes containing trisodium citrate dihydrate, and tissue samples were homogenized in PBS (protease inhibitors were added to PBS) on ice. Then, the samples were competed with a fixed amount of target on the solid phase supporter, where the target sites were biotinylated detection antibodies specific to the target. Subsequently, the HRP-conjugated streptavidin–biotin complex (SABC) was added to each well of the microplate and incubated. The enzyme–substrate reaction was initiated by the addition of TMB (3,3′,5,5′-tetramethylbenzidine) substrate solution, and the resulting color change was measured at a wavelength of 450 nm after terminating the reaction with a sulfuric acid solution.
Sperm motility
The caudal region of the epididymis from each mouse was placed in a centrifuge tube containing Hank’s medium. Subsequently, the caudal region was transferred to a fresh dish with Hank’s medium and punctured at multiple points along the longitudinal axis using a surgical dissection blade to release sperm into the medium. The dish was then incubated at 37 °C for 5 min to allow sperm dispersion. Sperm motility parameters were evaluated using a computer-assisted sperm analysis (CASA) system equipped with Sperm Vision HR software version 1.01 (Minitube, Ingersoll). Each sample was subjected to the analysis of 200 spermatozoa.
Sperm DNA integrity
Sperm DNA fragmentation was assessed using the Sperm Nucleus DNA Integrity Kit (Huakang Biomed) following the manufacturer’s guidelines. In brief, mouse semen samples were extracted from the epididymis and mixed with fluidized agarose in a tube. Approximately 30 μl of this mixture was deposited onto a pretreated slide and immediately covered with a coverslip. After a 4-min incubation at 4 °C, the coverslip was removed, and the slide underwent acid denaturation for 7 min, followed by 20 min of lysis. Subsequently, the slide was rinsed for 3 min with distilled water and sequentially dehydrated in 70%, 90% and 100% ethanol baths for 2 min each. After Wright’s staining, spermatozoa were counted to assess sperm DNA fragmentation using a bright-field microscope (Nikon).
Indirect calorimetry analyses
Continuous monitoring of O2 consumption, CO2 production, heat production and physical activity was conducted using a combined indirect calorimetry system (TSE phenoMaster, TSE Systems GmbH), as previously described15. The system was maintained at a constant environmental temperature of 22.0 ± 1.0 °C and followed a 12-h light/dark cycle (with lights off between 18:00 and 6:00). Individual housing in metabolic cages was provided for the mice, allowing unrestricted access to food and water.
Young and aged mice were acclimated to the metabolic cages for 48 h before data collection. All data were recorded during the next 48-h period in metabolic cages. Energy expenditure was determined by measuring heat production (kcal h−1 kg−1), O2 consumption (ml h−1 kg−1) and CO2 production (ml h−1 kg−1). Physical activity of each mouse was recorded over 48 h (counts per 48 h) using a multidimensional infrared light beam system. Bar graphs were generated based on the results obtained using the indirect calorimetry system, which mean light and dark phase results and all-time results.
Echocardiography
Cardiac function was evaluated using transthoracic echocardiography employing a Visual Sonics Vevo 3100 instrument equipped with a 30-mHz linear transducer probe (Fujifilm), as previously described62. Young and aged mice were induced with 2.5% isoflurane delivered in 1 L min−1 of compressed air, and anesthesia was sustained through an operative circuit nose GeroScience cone (Harvard Apparatus). Body temperature was monitored and maintained at 37 °C using a heating platform. Respiratory and heart rates were continuously monitored through a four-lead limb electrocardiogram, with isoflurane adjusted in 1 L min−1 compressed air to maintain a heart rate higher than 350 beats per minute (bpm). The mouse’s chest was shaved, and a layer of acoustic coupling gel was applied to the thorax. Subsequently, two-dimensional parasternal short-axis (SA) M-mode views of the left ventricle were acquired following optimization of gain settings, slight angulation and transducer rotation to enable imaging of both epicardial and endocardial surfaces. Particular care was taken during the procedure to prevent excessive thoracic pressure that could induce bradycardia. Cine loop analysis of cardiac cycles between respirations was conducted using VevoLab software to mitigate respiratory artifacts. Measurements of LV Vol;d, LV Vol;s, EF, FS and LV mass were obtained from a minimum of four cardiac cycles. Two observers, who were blinded to each other’s results, conducted all echocardiographic measurements.
Micro-CT scanning
In vivo micro-CT analysis was conducted to evaluate the microstructure of the proximal tibia and vertebrae in both young and aged mice. Anesthesia was induced using 2.5% isoflurane delivered in 1 L min−1 compressed air and maintained through an operative circuit nose cone provided by GeroScience. Respiratory and heart rates were continuously monitored using a four-lead limb electrocardiogram. Micro-CT images were acquired using a Hiscan XM Micro CT analyzer (Suzhou Hiscan Information Technology), following previously established protocols63. All scans were performed with an 80-kV tube voltage and a 100-A tube current, capturing images at a 25-µm resolution. A 0.5° rotation step was employed over a 360° angular range, with a 50-ms exposure per step. Image reconstruction was carried out using Hiscan Reconstruct software (version 3.0, Suzhou Hiscan Information Technology), and subsequent analysis of the original three-dimensional (3D) images of the proximal tibia and vertebrae was performed using Hiscan Analyzer software (version 3.0, Suzhou Hiscan Information Technology). From the CT scans, parameters including tTb.Th, Tb.Sp, Tb.N and BV/TV were calculated for each mouse. Two independent observers, who were blinded to each other’s results, conducted all measurements on the micro-CT images.
MRI scanning
Anesthesia was induced in both young and aged mice using 2% isoflurane and maintained at 1% throughout the MRI experiments. An oxygen generator with a flow rate of 2 L min−1, consisting of 90% oxygen and 10% isoflurane, was used and remained unchanged throughout the experiment. The mice were positioned in a horizontal bore 9.4T BioSpec 94/20 USR animal scanner (Bruker, Billerica). The body temperature and respiration rate of young and aged mice were continuously maintained using a warming pad and monitored using an MRI-compatible monitoring system. After scanning, young and aged mice were again transferred to the recovery room. T2WI and DTI protocols were used to ensure that images covered the volume between the olfactory bulb and the anterior part of cerebellum (field of view: 2.6 cm × 2.6 cm; thickness: 0.3 mm; 0.0703 × 0.0703 × 0.6 mm voxels; and 30 slices with no interslice gaps). Regions of interest (ROIs) were manually traced for the whole brain, hippocampus and cortex using the professional 3D medical image segment software ITK-SNAP 3.8.0 package (University of Pennsylvania) and based on previously reported methods64,65. The segmentation and volume calculations of MRI images were performed independently by two observers who remained blinded to each other’s results.
SA-β-gal staining
SA-β-gal staining was conducted using a commercial β-galactosidase staining detection kit (Solarbio)66. Cryosections were fixed with the supplied fixative solution for 15 min at room temperature. After fixation, the sections were immersed in a fresh β-gal staining solution in buffer and incubated at 37 °C overnight. Microscopic images of the stained sections were captured using a microscope (Nikon). The intensity of SA-β-gal staining in these images was quantified using ImageJ software (National Institutes of Health).
Immunohistochemistry of Ki67
Immunohistochemistry staining for Ki67 in the hippocampal sections was performed following the established protocol67. Hematoxylin staining was used to visualize cellular nuclei, which manifest as a blue hue, and the detection of Ki-67-positive staining relied upon the presence of discernible brown staining.
Quantification of AGE and ROS levels by ELISA
Quantification of AGE27 and ROS26 levels was performed using ELISA kits following the manufacturer’s instructions. In brief, samples were homogenized in cold PBS, and the total protein concentrations were determined using the BCA Kit (Thermo Fisher Scientific). The levels of AGEs were quantified using a commercially available ELISA kit (mouse AGE ELISA kit, FineTest Biotech) in accordance with the manufacturer’s protocol. This process involved the use of a multifunctional microplate reader (SpectraMax i3x, Molecular Devices). The content of ROS was measured using the same method and kit (mouse ROS ELISA kit, FineTest Biotech).
Lipofuscin staining (aldehyde fuchsine method)
Lipofuscin staining was conducted using a commercially available lipofuscin staining detection kit (Solarbio) in accordance with the manufacturer’s instructions68. The liver, kidney, heart and hippocampus were fixed in 4% (v/v) paraformaldehyde for 1 h. After dehydration and cryosectioning, tissue sections were stained in aldehyde fuchsine dyeing solution overnight at 37 °C. After this incubation, microscopic images of the sections were acquired using a microscope (Nikon).
Morris water maze test
The Morris water maze test was conducted in accordance with established protocols to evaluate spatial memory performance69. In brief, a circular water pool measuring 1.2 m in diameter and 50 cm in height was filled with water at a constant temperature of 24 °C. To facilitate accurate identification of the C57BL/6J mice, white food-grade stain was added to the pool. An imperceptible fixed platform, measuring 10 cm in diameter, was situated 1 cm below the water’s surface. The mice were trained to navigate and locate the concealed platform within the water pool. To achieve this, the mice were exercised to form a ‘spatial orientation map’ in the brain using different figure shapes and colors as visual stimuli from extra-maze cues along the sidewall of the swimming pool to accomplish this task.
During the spatial training phase (days 1–5), mice were given a 60-s time limit to locate the hidden platforms. The trial concluded automatically once a mouse successfully located the platform, climbed onto it within 60 s and remained on the platform for at least 5 s. Each day, the mice underwent four trials starting from different directions, with intervals of more than 20 min between trials. In case a mouse failed to find the platform within the allocated 60 s on the first day of training, gentle guidance was provided to lead it to the platform. The time taken by the mice to locate and ascend to the platform was documented as the escape latency, using a video camera for accuracy. After the spatial training phase, a probe trial was conducted on day 6 to evaluate memory consolidation. During this trial, the platform was removed from the pool, allowing the mice to swim freely for 60 s. The time spent by each mouse in the target quadrant, where the platform was originally positioned, as well as the number of times the mice crossed the location where the platform had been, were recorded by video and subsequently quantified.
During the acquisition training phase, aged mice consistently displayed prolonged escape latency and traversed longer paths in their quest to locate the concealed platform70. In contrast, young mice exhibited shorter escape latencies and covered shorter distances in comparison70. In the subsequent probe trial, young mice demonstrated a higher propensity to spend time in the target quadrant and made more frequent crossings over the original platform location, whereas their aged counterparts spent notably less time in the target quadrant and primarily navigated the tank without crossing over the platform’s initial position70. These observations collectively suggest compromised spatial learning and memory performance in the aged mice.
In this test, the variables were strictly controlled to ensure rigor and reproducibility of behavioral tests. The water temperature was kept constant, and appropriate lighting was provided. The experimental environment (for example, humidity, temperature, smell and sound) was kept constant in all behavioral sessions. The animals were consistently handled and tested by the same investigators throughout the entire duration of the Morris water maze tests. Moreover, the investigators conducting the behavioral experiments remained blinded to the group assignments. All experiments were conducted between 10:00 and 17:00.
Contextual fear conditioning test
The contextual fear conditioning test was conducted following established protocols71. In this test, mice were trained to associate the fear conditioning chamber with an aversive stimulus (mild foot shock; unconditioned stimulus (US)) for the assessment of contextual fear conditioning. Additionally, they were conditioned to associate tone cues (conditioned stimulus (CS)) with mild foot shocks to evaluate cued fear conditioning. The mice’s conditioned fear behavior was assessed by measuring freezing behavior.
On the training day (day 1), mice were placed in a fear conditioning chamber and allowed to acclimate to the environment for 2 min. After this, they received a 30-s tone (70 dB, 2 kHz) paired with a 2-s foot shock (0.6 mA). The CS–US pairing was repeated twice with a 2-min intermission. On the testing day (day 2), mice were placed in the same fear conditioning chamber for 3 min without any tone or foot shock to assess contextual memory. Freezing time was recorded and analyzed for this purpose. After a 2-h period, the mice were transferred to a new chamber with different odor, floor texture, chamber wall texture and chamber shape. They were given 2 min to acclimate, followed by a 30-s tone (70 dB, 2 kHz), and the freezing behavior was recorded and analyzed for 2 min to assess cued memory. The intensity of the foot shock was set at a level that was not painful but generated an unpleasant feeling for the mice. Freezing behavior was quantified and analyzed using a video tracking system and software (Taimeng).
Young and aged mice typically displayed similar levels of baseline freezing behavior during the training phase. However, during contextual memory testing, aged mice consistently exhibited lower levels of freezing compared to young mice72. These observations suggest the presence of non-spatial deficits in learning and memory in aged mice.
Exhaustive running test
A treadmill running test was used to measure the endurance exercise capacity and was performed as described previously69. In brief, acclimatization to the treadmill was carried out over a period of 2 d before the experimental phase. This involved a 10-min run at 10 m min−1, followed by a 2-min run at 20 m min−1.
On the day of the exhaustive exercise, the mice were subjected to a progressive running protocol on a fixed 10° slope. This protocol consisted of running at 10 m min−1 for 15 min, followed by 13 m min−1 for 3 min, 16 m min−1 for 3 min, 19 m min−1 for 3 min and 22 m min−1 for 3 min, after which they ran at 25 m min−1 until exhaustion. Forced running was induced by positioning an electric grid with a voltage of 35 V at the end of the treadmill. Exhaustion was defined as the mice remaining on the shock grid for a continuous period of 5 s. The time to exhaustion was recorded. All animals were exercised at the same time to minimize diurnal effects. The animals were consistently handled and tested by the same investigators throughout the entire duration of the exhaustive running tests. Moreover, the investigators conducting the behavioral experiments remained blinded to the group assignments. All experiments were conducted between 10:00 and 17:00.
sEV labeling and tracking
sEVs were labeled using PKH26 fluorescence kits following the manufacturer’s guidelines. In brief, sEVs suspended in PBS were mixed with PKH26 dye. After ultracentrifugation at 120,000g for 70 min, the supernatant containing the unbound dye was discarded, and the pellet was resuspended in PBS. Labeled sEVs were then intravenously injected into wild-type C57BL/6J mice. On day 2 after administration, the mice were anesthetized and transcardially perfused with ice-cold 0.9% saline to eliminate any remaining blood-derived sEVs in tissues. Hippocampus and muscle were excised, and frozen sections were analyzed using fluorescence confocal microscopy to visualize the red fluorescent signals.
Protein reduction, alkylation, digestion and iTRAQ labeling
Protein extraction, protein concentration measurement and iTRAQ labeling were performed as previously described73. In brief, 100 µg of protein isolated from eight different organs/tissues (including heart, liver, spleen, lung, kidney, hippocampus, muscle and testis) was used for iTRAQ experiments. After TCEP (AB SCIEX) reduction, MMTS (AB SCIEX) alkylation and trypsin (Promega) digestion, the resulting peptides from each organ/tissue were collected and subjected to iTRAQ labeling using an iTRAQ Reagent 8-Plex Multiplex Kit (AB SCIEX). In one individual iTRAQ experiment, four biological replicate samples from PBS-injected aged C57BL/6J male mice were labeled with iTRAQ tags 113, 114, 115 and 116, and four samples from young-sEV-injected aged mice were labeled with iTRAQ tags 117, 118, 119 and 121. For each iTRAQ experiment, all labeled peptides were finally separated into 16 samples, desalted and concentrated for liquid chromatography with tandem mass spectrometry (LC–MS/MS) analysis, as previously described73.
LC–MS/MS and database searching
MS data were acquired using a TripleTOF 5600+ System (AB SCIEX), as previously described73. The original MS/MS data were submitted to ProteinPilot version 4.5 (AB SCIEX) for database searching by searching against Mus musculus sequences in the UniProt database concatenated with the reverse decoy database (4 March 2021, containing 55,366 sequences; http://www.UniProt.org/proteomes/UP000000589). The searching parameters were set as described in a previous study73. The MS proteomics data have been deposited to the ProteomeXchange Consortium74 via the PRIDE75 partner repository with project accession code PXD030350.
Bioinformatics analysis
Intensive bioinformatics analyses were conducted with R, RStudio, GSEA and other requisite packages and online tools.
To get an overview of quantitative proteomics data from all eight groups, both the identified proteins and the quantified proteins were counted for each group, and UMAP analysis was performed using the R package ‘umap’. Moreover, a whole-proteome heatmap was also generated by the R package clusterProfiler76 to determine the homogeneity in four samples injected with PBS in all eight organs/tissues and four samples injected with young sEVs in the same iTRAQ experiment as a quality control of biological replicates.
For in-depth functional class scoring, GSEA (http://www.gsea-msigdb.org) was performed using the R package clusterProfiler76 and GSEA software77. The significantly upregulated or downregulated GO terms between four samples injected with PBS and four samples injected with young sEVs in all eight tissues or organs were retained for further analysis. The GO terms of GSEA were clustered with the ‘binary cut’ algorithm using the R package simplifyEnrichment to further analyze the GSEA results. In the process of ‘binary cut’ clustering, difference score, cluster numbers and other parameters were set following the R package documentation78. Heatmaps with word cloud annotations were generated to summarize the functions of each enriched GO cluster.
Protein extraction and western blots
For cellular protein isolation, cells were rinsed with PBS (pH 7.4) and subsequently lysed in RIPA lysis buffer supplemented with PMSF (1:100, Beyotime Biotechnology) and a phosphatase inhibitor cocktail (1:100, Thermo Fisher Scientific) at 4 °C for 30 min. For tissue protein isolation, samples mixed with RIPA, PMSF and phosphatase inhibitor cocktail were homogenized using a grinder at 60 Hz for 60 s, followed by lysis at 4 °C for 30 min. Subsequently, cell lysates and tissue homogenates were centrifuged at 12,000g and 4 °C for 10 min. The resulting supernatant was collected, and protein concentrations were determined using the BCA Protein Assay Kit (Thermo Fisher Scientific).
Samples containing equivalent amounts of protein (40–80 μg) were loaded and separated on 10–12.5% SDS-PAGE gels and subsequently transferred to PVDF membranes (Millipore). These membranes were blocked for 1 h at room temperature with 5% non-fat dry milk in TBS containing 0.1% Tween 20. After blocking, the membranes were incubated overnight at 4 °C with the appropriate primary antibodies. After three washes with 1× TBST for 10 min each, secondary antibodies were incubated for 1 h at room temperature, followed by an additional three 10-min washes with 1× TBST. The membranes were treated with ECL substrate (Thermo Fisher Scientific) following the manufacturer’s instructions, and protein levels were visualized using the Tanon 5200 multi-detection system (Tanon). Data were quantified using ImageJ software, and the relative protein levels were normalized to the β-actin level. Additional information on primary antibodies is available in Supplementary Table 4.
RNA isolation and quantitative RT–PCR assay
Total RNA was extracted from cell and tissue samples using TRIzol reagent (Invitrogen) following previously described methods69.
For mRNA expression analysis, cDNAs were synthesized from 1 μg of total RNA using the AMV reverse transcriptase (TaKaRa) and oligo dT primers (TaKaRa). The reverse transcription reaction was carried out at 16 °C for 5 min, 42 °C for 60 min and 85 °C for 5 min. Quantitative RT–PCR was performed on an Applied Biosystems 7300 Sequence Detection System using a SYBR Green PCR kit. The PCR reactions underwent initial denaturation at 95 °C for 10 min and were then amplified for 40 cycles, with denaturation at 95 °C for 15 s, annealing at 60 °C for 30 s and extension at 72 °C for 30 s. All reactions were run in triplicate.
For miRNA expression analysis, cDNAs were synthesized from 500 ng of total RNA using the miRNA 1st Strand cDNA Synthesis Kit (Vazyme) with the stem-loop method. The reverse transcription reaction was carried out at 16 °C for 30 min, 42 °C for 30 min and 85 °C for 5 min. Quantitative RT–PCR was performed on the Applied Biosystems 7300 Sequence Detection System using miRNA Universal SYBR qPCR Master Mix (Vazyme) in a 96-well optical plate. The thermal cycling conditions included an initial denaturation step at 95 °C for 10 min, followed by 40 cycles at 95 °C for 15 s and 60 °C for 1 min. All reactions were run in triplicate.
After completion of the reactions, the CT values were determined using fixed threshold settings, and the mean CT was calculated from triplicate PCR measurements. The 2−ΔΔCT method was employed to determine the relative expression of RNAs, with β-actin mRNA serving as the internal control for quantification of mRNAs and miR-16 serving as the internal control for quantification of miRNAs. The primer sequences used are provided in Supplementary Table 5.
Measurement of ATP
The rate of ATP synthesis was determined using an ATP Determination Kit (Biovision) following the manufacturer’s instructions. This assay employs luciferase to facilitate the production of light from ATP and luciferin, which is subsequently measured using a luminometer. In brief, samples were homogenized with a nuclear releasing buffer for 5 min at room temperature. After this, lysate was added to the ATP monitoring enzyme for the calculation of ATP synthesis within 1 min using a luminometer. Protein concentrations were determined using the BCA Protein Assay Kit (Thermo Fisher Scientific) and applied for data normalization.
Measurement of relative mtDNA content
mtDNA copy number was quantified using quantitative PCR, following a previously described protocol31. In brief, total DNA was extracted from tissues and cells using a DNA extraction kit (Biomed), in accordance with the manufacturer’s instructions79. A total of 10 ng of DNA was used for quantitative PCR analysis. The relative mtDNA copy number was determined by measuring the amplification of mitochondrially encoded genes, including cytochrome c oxidase I (MT-CO1), NADH dehydrogenase 1 (MT-ND1), cytochrome c oxidase III (MT-CO3) and the D-loop region. These measurements were normalized to β2-microglobulin (β2-MG). The primer sequences used in the mtDNA assay are provided in Supplementary Table 5.
Measurement of mitochondrial complex V activity
The activity of mitochondrial complex V was assessed using a mitochondrial respiratory chain complex V activity detection kit (Yifeixue Bio Tech), in accordance with the manufacturer’s instructions. In brief, tissues or cells were homogenized or ground, and mitochondria were extracted. Sonication was performed on ice using an ultrasonic cell disruptor (Sonics) at 200 W for 3 s, repeated 30 times at 10-s intervals. After a 30-min enzymatic reaction and a 10-min chromogenic reaction, the absorbance was measured using a microplate reader (Molecular Devices).
SDHA immunofluorescence staining and SDH staining
SDHA immunofluorescence staining and SDH staining were performed following previously established protocols with slight modifications80. Muscle tissues were fixed with 4% paraformaldehyde in PBS (pH = 7.4) for 48 h at 4 °C. The fixed tissues were then dehydrated using a 30% sucrose solution. Cryosectioning was performed using a cryomicrotome (Leica). The sections were subsequently incubated overnight with a mouse anti-SDHA primary antibody (1:500, Abcam, ab814715), followed by staining with fluorescent dye-conjugated secondary antibodies (Life Technologies) for 1 h at room temperature. Finally, the sections were imaged using a confocal microscope (Nikon).
For SDH staining, mouse muscle samples were immediately immersed in OCT solution and frozen at −80 °C. Frozen sections with a thickness of 7 μm were prepared using a cryostat (Leica). These sections were then incubated with SDH reaction reagent at 37 °C for 40 min. After the enzymatic reaction, the reaction was halted by a rapid immersion in ddH2O and subsequent air drying for 10 min in the absence of light. Capturing images of the sections was conducted using a microscope (Nikon). The staining intensity of SDH in the images was quantified using ImageJ software.
TEM analysis of the mitochondrial ultrastructure
Hippocampal and muscle samples were trimmed to 2 mm3 and fixed in 2.5% glutaraldehyde overnight at 4 °C. Subsequently, the samples underwent treatment with 1% osmium tetroxide and dehydration through a gradient of ethanol solutions. After staining with 1% uranylacetate and dehydration in acetone, samples were sliced into ultra-thin sections (80 nm) and loaded on electron microscopy grids. Images were acquired using a Hitachi TEM system (Hitachi, HT7700) operating at an acceleration voltage of 80.0 kV. The number of mitochondria was calculated in three fields for each sample section at ×20,000 magnification. The ultrastructure of mitochondria was captured at ×70,000 magnification and was identified as damaged if samples had disrupted membranes, cristae depletion, matrix dissolution and vacuolization.
Cell culture
The mouse embryonic neural stem cell line NE-4C and mouse myoblast cell line C2C12 were obtained from the Shanghai Institute of Cell Biology, Chinese Academy of Sciences. NE-4C cells were cultured in complete mouse neural stem cell medium (Procell) at 37 °C with 5% CO2. To facilitate NE-4C cell adhesion and growth, 10 μg ml−1 poly-l-lysine was applied to the flask for at least 2 h before cell passage and cultivation. Furthermore, NE-4C cells were passaged using 0.05% trypsin instead of 0.25% trypsin. C2C12 cells were maintained in high-glucose (4.5 g L−1) DMEM (Gibco) supplemented with 10% FBS (Gibco, 10099141C) and cultured at 37 °C with 5% CO2.
Upon reaching 70% confluence, NE-4C cells were subjected to neuronal differentiation by supplementing the culture medium with 10 μM retinoic acid. After a 6-d differentiation period, NE-4C cells were either co-cultured with sEVs or transfected with miRNA mimics and antisenses. Similarly, C2C12 cells were induced to undergo the myoblast–myotube transition by switching the growth medium to differentiation medium (DMEM supplemented with 2% horse serum) when they reached 90% confluence. After 1 d, C2C12 cells were either co-cultured with sEVs or transfected with miRNA mimics and antisenses.
Induction of cell senescence with high dose of H2O2
Cell senescence was induced following a previously described protocol81. On the first day, cells were cultured in T175 flasks and assessed under an inverted microscope (Nikon) to ensure that they reached approximately 50% confluence. If the cells had reached the desired confluence, the complete growth media were replaced with pre-warmed basal media containing 1.0 mM H2O2. The cells were then incubated in a cell culture incubator at 37 °C for 1 h. Afterwards, the basal media were changed back to complete growth media, and the cells were further incubated at 37 °C for the next 23 h. The next day, the cells were examined under an inverted microscope to confirm their viability and health. Subsequently, the complete growth media were replaced with pre-warmed basal media containing 750 μM H2O2. The cells were once again incubated in a cell culture incubator at 37 °C for 1 h. After that, the basal media were changed back to complete growth media, and the cells were incubated at 37 °C for another 23 h. The aforementioned steps were repeated four times until day 6 to induce cell senescence.
Co-culture with sEVs and transfection with synthetic RNAs
For the co-culture experiments with sEVs, sEVs (50 μg of total protein content) were incubated with 1 × 106 NE-4C or C2C12 cells and cultured in a humidified atmosphere with 5% CO2 at 37 °C. After 24 h, total RNA and protein were extracted from the cells. For cell transfection experiments, 50 pmol of miRNA mimics, antisense miRNAs or corresponding scrambled RNA (scrRNA) (GenePharma) were transfected into NE-4C or C2C12 cells using Lipofectamine 2000 (Invitrogen) following the manufacturer’s instructions. Total RNA and protein were extracted 48 h after transfection. The sequences of the synthetic miRNA mimics and antisenses can be found in Supplementary Table 6.
EdU incorporation assay
An EdU incorporation assay was conducted following the previously described protocol with slight modifications82. NE-4C or C2C12 cells were seeded in 48-well plates after co-culture with sEVs or transfection with miRNA mimics or antisenses. When cells reached 80% confluence, they were treated with EdU reagent (RiboBio, C10310-1) to assess cell proliferation according to the manufacturer’s instructions. Cell nuclei were stained using DAPI (Beyotime Biotechnology), and the stained cells were visualized using a BX51 fluorescence microscope (Olympus).
Measurement of OCR
OCR was analyzed using an XF96 Analyzer (Seahorse Bioscience). In brief, 2 × 104 cells per well were plated in Seahorse XF96 plates in 200 μl of DMEM supplemented with 1 mM sodium pyruvate, 2 mM glutamine and 10 mM glucose. The plates were incubated for 1 h at 37 °C in a humidified incubator without CO2. After assessing the OCR at baseline, the following compounds were sequentially added to the analyzer, and the change in OCR was measured: (1) oligomycin (1.5 μM), which blocks ATP synthase and reduces OCR, representing mitochondrial respiration associated with cellular ATP production (ATP-coupled OCR); (2) carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone (FCCP, 0.5 μM), a mitochondrial uncoupler that enables maximal rates of electron transport and measures the maximal respiratory capacity (maximal OCR); and (3) a combination of rotenone and antimycin A (0.5 μM), which blocks respiratory electron flux through complexes I and III and inhibits mitochondrial respiration. All OCR measurements were normalized to the cell number in each well.
Small RNA deep sequencing
Small RNA deep sequencing was conducted by BGI (Shenzhen) using a TruSeq Small RNA Sample Prep Kit (Illumina) for the construction of small RNA libraries. After the quality validation of these libraries, raw data were generated using the Illumina HiSeq 4000 platform. The clean reads were obtained through data filtration. Sequences corresponding to mature miRNAs and their precursors were extracted from miRBase version 21. Subsequently, the clean reads were aligned to reference sequences and annotated using the Bowtie algorithm. Only candidates with no more than one mismatch and not exceeding two shifts were considered as miRNA matches. Statistical analysis was conducted using Student’s t-test, with statistical significance defined as uncorrected P < 0.05 and |fold change| > 2. The average expression threshold was set at more than 50 reads.
Literature mining of age-related circulating miRNAs
Age-related circulating miRNAs were identified through a PubMed-based literature mining approach. In brief, we conducted searches using several keywords, such as ‘plasma miRNA AND aging’, ‘serum miRNA AND aging’, ‘plasma miRNA AND metabolism’ and ‘serum miRNA AND metabolism’, on PubMed, applying the ‘Original article’ filter. We extracted all miRNAs reported in the literature and consolidated them after removing duplicates. A total of 458 miRNAs were identified in the published studies and subjected to further manual classification analysis. miRNAs were included if the studies focused on mouse or human miRNAs, if miRNA levels were measured using RNA deep sequencing or quantitative RT–PCR and if miRNAs were examined in at least three published papers. miRNAs were excluded if contradictory reports existed regarding their alterations in aged plasma (serum) compared to young plasma (serum). Ultimately, we identified 90 age-related circulating miRNAs, comprising 53 that exhibited increased levels and 37 that showed decreased levels with advancing age.
Statistics and reproducibility
Statistical analyses were conducted using GraphPad Prism version 8 (GraphPad Software) or the open-source statistical package R. Significance was determined using two-sided Student’s t-test or one-way ANOVA followed by Dunnett’s multiple comparison test. Significance was assumed at *P < 0.05, **P < 0.01 and ***P < 0.005. Details on the statistics used can be found in the figure legends. No animals or data points were omitted from the analysis for any particular reason. The data are presented as the means ± s.e.m. n represents the number of samples used in the experiments. All micrographs and blots were representative of three independent experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.