Structure determination
To obtain a stable and homogenous human SVCT1 (hSVCT1) sample suitable for cryo-EM analysis, we first screened solubilization conditions for hSVCT1. The C-terminally GFP-fused hSVCT1, with functional properties equivalent to those reported previously13 (Fig. 1a, b), was transiently expressed in HEK293T adherent cells and solubilized with various detergents, and then the degrees of stability and monodispersity were examined by fluorescent-detection size-exclusion chromatography (FSEC)25 based on the GFP fluorescence. hSVCT1 exhibited two main elution peaks (Supplementary Fig. 1a), likely corresponding to different oligomeric forms, especially monomers and dimers, as expected from previous research19,20,26. Since the importance of dimer formation for the transport activity has been reported for the bacterial homolog UraA20 and fungal homolog UapA21, and larger molecular weights and more symmetrical properties of the target are generally advantageous in cryo-EM single particle analysis, we focused on the first peak possibly corresponding to the dimeric form, which predominantly appeared in the lauryl maltose neopentyl glycol supplemented with cholesteryl hemisuccinate (LMNG-CHS)-solubilized conditions. We expressed hSVCT1-GFP in HEK293S GnTI− cells and purified it with GFP nanobody-immobilized resins, followed by size-exclusion chromatography in the presence of Na+ (Supplementary Fig. 1b, c). Purified hSVCT1 preparations with or without vitamin C (sodium ascorbate) were subjected to cryo-EM single particle analysis, and we determined the inward-open structures of hSVCT1 in the substrate-bound and substrate-free states at overall resolutions of 2.5 and 2.6 Å, respectively (Fig. 1c and Supplementary Figs. 2 and 3). The two structures have no significant differences except for the substrate binding, and are aligned with a root mean square deviation (RMSD) of 0.28 Å for 1,024 Cα atoms. Therefore, with regard to the inward-open conformation, we describe the structural features of the well-resolved substrate-bound state for simplicity, unless otherwise specified. We also prepared the hSVCT1 sample purified in the absence of Na+ (Supplementary Fig. 1d–f), and obtained its 2.8 Å resolution occluded structure by cryo-EM analysis (Fig. 1d and Supplementary Fig. 4). The details and comparisons of this novel structure will be discussed in a later section.

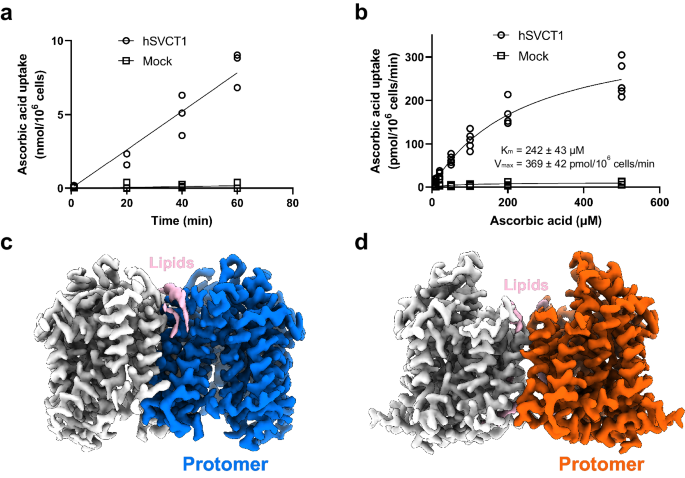
a Time-dependent ascorbic acid transport into HEK293T cells by hSVCT1 (n = 3 biological replicates). b Km curve for hSVCT1-derived ascorbic acid uptake into HEK293T cells (n = 5 for hSVCT1, n = 3 for mock). Source data are provided as a Source Data file. c Cryo-EM map of the hSVCT1 homodimer in the inward-open state, with densities corresponding to each protomer and lipids colored blue, gray, and pink, respectively. d Cryo-EM map of the hSVCT1 homodimer in the substrate-free occluded state, with densities corresponding to each protomer and lipids colored orange, gray, and pink, respectively.
Overall structure of hSVCT1 in the inward-open state
hSVCT1 forms a homodimer, with each protomer consisting of 14 transmembrane helices (TMs) organized into a “core domain” (TMs 1–4 and 8–11) and a “gate domain” (TMs 5–7 and 12–14) (Fig. 2a, c–e and Supplementary Fig. 5). This protein fold of the protomer is referred to as the “UraA fold”, and is conserved among the SLC4, 23 and 26 transporter families19,20,21,27,28,29,30. The surface model showed that it adopts an inward-open state with a conserved substrate binding pocket accessible from the cytoplasmic side (Fig. 2b). A cleft is formed at the interface of the core and gate domains, and the substrate binding pocket is located at the end of the cleft, backed by a pair of short antiparallel β-strands between two half-helices of TMs 3 and 10 (Fig. 2e). Considering the similarity of the two obtained inward-open structures, we assumed that the substrate-bound and substrate-free structures correspond to the states before and after substrate release, respectively.

a Overall structure of inward-open hSVCT1. The core and gate domains of a protomer are colored cyan and blue, respectively. b Electrostatic surface representation of inward-open hSVCT1 sliced at the transport pathway. Vitamin C bound within the substrate binding pocket is shown as yellow sticks. c Protomer viewed from the extracellular side, with each TM numbered. d Gate domain and e core domain viewed from the other protomer.
The formation of the hSVCT1 dimer is predominantly achieved by interactions between the TM regions in a similar manner to its homologs20,21, where the gate domains face each other with an extensive interface of ~2,000 Å2, partly mediated by lipids (Fig. 1c and Supplementary Fig. 6). This is in stark contrast to the SLC4 and 26 transporters, where the gate domains form either a smaller or no interface27,29. The SLC4 and 26 transporters have a large cytosolic domain that connects two protomers29,31,32, while hSVCT1 has a small structural component outside the plasma membrane, suggesting that the absence of a cytosolic domain in the SLC23 family, including SVCT1, facilitates the formation of a predominant dimeric interface between the TM domains. An N-linked glycan is present at Asn144 on the extracellular loop connecting TMs 3 and 4 (Supplementary Fig. 6g), where N-glycosylation is reportedly important for membrane targeting and substrate transport33. In addition, S-acylation is observed at Cys565, located in the protruding loop region at the border between the plasma membrane and cytoplasm (Supplementary Fig. 6h). Although its function has not been reported so far, the S-acylation of this residue, with its attached fatty acid chain inserted within the membrane, may contribute to protein stability or membrane trafficking34.
Vitamin C recognition mechanism
We identified a vitamin C molecule bound at the conserved substrate binding site inside the protomer. Our density map is clear enough to reliably determine its orientation (Fig. 3a and Supplementary Fig. 7a). Several non-protein densities are observed at the pocket in addition to vitamin C, and we modeled two sodium ions and three water molecules according to the coordination of the surrounding residues (Fig. 3b–d). The presence of two sodium ions together with one vitamin C molecule is consistent with previous reports that SVCTs transport sodium ions and vitamin C in a 2:1 stoichiometry11,12,35. As the substrate binding pocket is potentially negatively charged (Fig. 2b), sodium ions are required to neutralize the negative charge to facilitate vitamin C binding, confirming the proposed mechanism reported for the mouse SVCT1 structure24. Additionally, TMs 3 and 10 sandwich vitamin C, and their dipole moments electrostatically stabilize the bound substrate, as observed in the Cl– binding of Prestin30. The first sodium ion, Na1, is surrounded by TMs 8 and 10, as well as vitamin C. Na1 interacts with the sidechains of Glu334, Asp338, and Ser383, the backbone carbonyl of Ser381, and an oxygen atom of vitamin C (Fig. 3b). The location and coordination manner of Na1 are generally consistent with those in mouse SVCT124 and similar to the sodium ion binding mode of the SLC4 transporter NDCBE36. A notable feature in our structure is the direct interaction between Glu334 and vitamin C, which emphasizes the importance of the conserved glutamate residue in substrate recognition. A comparison between the substrate-bound and substrate-free structures revealed that the sidechain of Ser383 is flipped after releasing the substrate and directly interacts with the sidechain of Asp338, without mediation by a sodium ion, suggesting that substrate release triggers the reorganization of the substrate binding pocket, and possibly induces the subsequent conformational change (Supplementary Fig. 7b). The second sodium ion, Na2, is located opposite Na1 across the vitamin C, where strong density extending from the substrate is observed (Fig. 3a, c and Supplementary Fig. 7a). Na2 is stabilized by a cation-π interaction with the aromatic sidechain of Phe112, as well as by polar interactions with Ser110 and vitamin C. Notably, the position of Na2 in our structure is different from that in the mouse SVCT1 structure, in which Na2 is surrounded by TMs 3 and 8 on the same side as Na1 (ref. 24; Supplementary Fig. 8). Although non-protein density is also observed at that position in our density map, we modeled a water molecule based on the hydrogen bonds from vitamin C and the backbone amide of Ala113 (Fig. 3d). Under neutral pH conditions, vitamin C exists as a monovalent anion in resonance between two forms, with a negative charge on the oxygen atom covalently bonded to either the C1 or C3 carbon2,37 (Supplementary Fig. 8). In our structure, two sodium ions are located beside these potentially negatively charged oxygen atoms, thus confirming the stable coordination manner of the positively charged ions. Together with the two sodium ions and three water molecules, the formation of a broad hydrogen bond network around the vitamin C is likely to play a role in stabilizing the substrate binding. Another interaction is the polar interaction between Thr59 and the oxygen atom of the vitamin C’s 1,2-dihydroxyethyl group (referred to as the “tail” for simplicity; Fig. 3c and Supplementary Fig. 9). While the lactone ring of vitamin C is strongly stabilized by interactions with the core domain through two sodium ions and water molecules, the tail forms relatively few interactions with the protein chain and neighboring small molecules. This biased distribution of interactions with the substrate possibly contributes to the incomplete selectivity for vitamin C and its stereoisomer, where l-ascorbic acid is favorably transported and d-isoascorbic acid, which only differs in the orientation of the tail, is transported less efficiently but in detectable amounts10,38.

a Overall view of the substrate binding pocket. The green mesh represents the Fo–Fc omit map of vitamin C, two sodium ions, and three water molecules contoured at 3.0σ (where σ is the standard deviation within the mask). b–d Close-up views and interaction manners of each ligand. e Substrate uptake assays for point mutants. Values are mean of n = 3 biological replicates ± s.e.m. and compared relative to the wild-type uptake rate. Source data are provided as a Source Data file.
To examine the functional importance of the residues comprising the substrate binding pocket, we measured the transport activities of hSVCT1 mutants by a cellular ascorbic acid uptake assay, based on its reducing ability (Fig. 3e). The results demonstrated that the alanine mutations of the residues involved in the coordination of two sodium ions significantly decreased the transport activity, confirming the sodium-dependent binding manner of the substrate suggested from the structure. The mutation of Thr59, which interacts with the tail of the vitamin C molecule, caused a partial impairment of substrate uptake, consistent with the proposed weak recognition of this region. Although we did not detect any direct interactions between the substrate and residues such as Ser382 and Pro384, mutants of these residues exhibited rather decreased transport activities. These residues may have a role in substrate affinity by forming and stabilizing the hydrophilic pocket.
hSVCT1 structure in the substrate-free occluded state
To elucidate the molecular mechanism of the overall transport cycle, other conformations of the hSVCT1 structure must be solved. Focusing on the sodium-dependent transport by SVCTs, we assumed that hSVCT1 would adopt different transporting conformations under sodium-free conditions. After confirming that hSVCT1 exhibits a similar and stable oligomerization profile with the substitution of sodium with potassium in all buffers, we purified hSVCT1 dimers using the same procedure as for the sodium-containing conditions, except for the switching of cations (Supplementary Fig. 1d–f). The cryo-EM analysis, performed using the same procedure as for the sodium-containing conditions, resulted in two major 3D reconstruction classes (classes 1 and 2; Supplementary Fig. 4). Whereas class 1 showed a similar conformation to that of the inward-open states under sodium-containing conditions, class 2 showed a different conformation, whose particles resulted in a final refinement at 2.8 Å resolution (Figs. 1d and 4a and Supplementary Fig. 4). The substrate binding pocket in this map is vacant, and sequestered from the solvent mainly by the sidechains of Phe112, Glu334, and Ser383, suggesting that this structure represents a substrate-free occluded state (Fig. 4b, c). This substrate-free occluded structure reveals an unprecedented conformational state for the NAT family, representing a structure between the substrate-free inward-open and outward-open states. The occluded substrate binding pocket may be too small to accommodate a vitamin C molecule, indicating its different structural properties from the substrate-bound occluded structure previously reported for UraA20 (Supplementary Fig. 10). For the inward open-like class under the potassium conditions (class 1), the final reconstruction resulted in a density map at a resolution of 3.5 Å (Supplementary Figs. 4 and 11). The intracellular region of the core domain, especially TM10, is disordered in this structure, suggesting the flexibility of this region. This class possibly corresponds to an intermediate state during the transition from the inward-open state to the substrate-free occluded state, indicating the conformational equilibrium between two distinct states.

a Overall structure of substrate-free occluded hSVCT1. The core and gate domains of a protomer are colored yellow and orange, respectively. b Electrostatic surface representation of substrate-free occluded hSVCT1 sliced at the substrate binding pocket. c Close-up view of the occluded pocket. d, e Comparison of the core domains between inward-open and substrate-free occluded hSVCT1, aligned by the protomeric core domain. (d) Overall alignment of the core domain and (e) highlights of TMs 3 and 10. f Comparison of the gate domain aligned by the core domain of the same protomer. g–h Comparison of the residues comprising (g) the substrate binding pocket and (h) the extracellular gate.
A superimposition of the core domains of the inward-open and substrate-free occluded structures revealed a good alignment with an RMSD of 1.32 Å for 308 Cα atoms with no significant differences, except for the notable upward shift of TM10 towards the extracellular side from the inward-open to substrate-free occluded state (Fig. 4d, e). The up-and-down movements of the core domain, especially for TMs 3 and 10, were reported in previous studies of SLC proteins structurally related to NAT transporters29,30,32,39,40, and are referred to as elevator movements. UapA21 and mouse SVCT124 also adopt this elevator mechanism through the transition from the substrate-bound occluded to inward-open states, with TMs 3 and 10 shifting downward to the intracellular side. The movement captured here can also be regarded as an example of elevator-like movement, but a notable difference from the previous reports is the additional horizontal shift of TM10 toward the gate domain, which occludes the transport pathway and prevents access to the interior of the protein (Fig. 4e). Flexibility of TM10 observed in the intermediate structure also supports the large movement of this region (Supplementary Figs. 5c and 11). The residues comprising the substrate binding pocket, such as Ser382, Ser383, and Pro384, are shifted to the extracellular side along with the movement (Fig. 4g). The sidechains of Glu334 and Ser383, which are responsible for the coordination of Na1 in the substrate-bound inward-open state, are in close contact and directly interact without a sodium ion in this occluded state, as Asp338 and Ser383 in the substrate-free inward-open state, contributing to the formation of the intracellular gate (Fig. 4c, g and Supplementary Fig. 7b). Ser382, whose sidechain is not involved in the substrate recognition, is flipped and oriented towards the substrate binding pocket after the conformational change, suggesting the importance of this residue for the occlusion of the pocket after substrate release (Fig. 4g).
Superimposition of the protomers of the inward-open and substrate-free occluded structures relative to the core domain revealed that the entire gate domain tilts toward the cytoplasmic side of the core domain in the same protomer, with TMs 6 and 13 outwardly rotated in the transition from the inward-open state to the substrate-free occluded state (Fig. 4f). This tilting motion of the gate domain is likely related to the rocking bundle-like mechanism proposed in previous studies of UraA monomer and dimer structures19,20. We observed the flipping of the aromatic sidechains of Phe112 in the core domain and Phe436 in the gate domain through the conformational change (Fig. 4g, h). In the inward-open state, Phe112 forms a π-π stacking interaction to constitute the extracellular gate, while Phe112 also coordinates Na2 by a cation-π interaction on the opposite side of the phenyl group (Fig. 3c). In the substrate-free occluded state without a sodium ion, however, the stacking is disrupted due to the flipping of both phenylalanine residues and the extracellular gate is formed only by Phe112, indicating the fragility of the gate in this occluded state (Fig. 4c, h). Thus, this state possibly represents a state just prior to switching to the outward-open state and importing the next substrate.
Dimer formation is requisite for transport function
A comparison of the dimer interfaces between the inward-open and substrate-free occluded structures revealed an extensive conformational change (Fig. 5a, b and Supplementary Movie 1). In the inward-open state, a stable dimer interface is formed mainly by TMs 6, 12, and 13 of the gate domains, with TM13 inserted into a cleft on the intracellular side formed by TMs 6’ and 12’ of the other protomer (apostrophe indicates the adjacent protomer; Fig. 5a). However, in the substrate-free occluded state, the gate domains are reorganized to form a different dimer interface, in which TM13 has left the cleft and become exposed to the plasma membrane (Fig. 5b). Detailed observations of the dimer interfaces of the two states revealed the rearrangements of hydrogen bonds and the state-specific hydrophobic interactions between protomers. As for hydrogen bonds, Asn453’ on TM13’ forms a bipartite hydrogen-bond with Gln448 on TM12 on the intracellular side in the inward-open structure (Fig. 5c), while Asn453’ on TM13’ hydrogen-bonds with Asn230 on TM6 in the occluded structure, to stabilize the dimer formation in each state (Fig. 5e). As for hydrophobic interactions, multiple phenylalanine sidechains on TM13 are inserted into the cleft between TM6’ and TM12’ to form the unique dimer interface in the inward-open state, accompanied by a lipid molecule (Fig. 5d). In the occluded state, Trp211 on TM6, which faced outwardly from the protein in the inward-open state, has turned inside and interacts with Phe200’ on TM5’ to stabilize the formation of a differently organized dimer interface together with an LMNG molecule (Fig. 5f). In addition, Leu223 on TM6 is located adjacent to another LMNG molecule at the dimer interface, coordinating the close packing of the interface in the occluded structure (Fig. 5e).

a–b Representations of dimer interfaces in (a) inward-open state and (b) substrate-free occluded state, viewed from the intracellular side. c–f Close-up views of the residues responsible for dimer formation in (c–d) inward-open state and (e–f) substrate-free occluded state. g Evaluation of oligomeric population of dimer interface mutants by FSEC, detected by GFP fluorescence. h Substrate uptake assay for dimer interface mutants. Values are mean of n = 3 biological replicates ± s.e.m. compared relative to the wild-type uptake rate. Source data are provided as a Source Data file.
To investigate the importance of these residues in the distinct dimer interactions and the transport activity of hSVCT1, we introduced mutations of these residues to diminish the interactions between protomers or overlap with the lipid molecule. We examined the ratio of monomeric or dimeric hSVCT1 population by FSEC, and found that the FSEC peak partially shifted from the dimeric to monomeric position for all mutants (Fig. 5g). Notably, mutants of hydrogen-bonded residue pairs exhibited a gradual decrease in dimer formation as more mutations were introduced for both the Gln448/Asn453 and Asn230/Asn453 pairs, supporting the importance of these residues for stabilizing the dimerization and the existence of two states under physiological conditions. We tested the transport activities of these mutants, and found that vitamin C transport decreased for all mutants in accordance with the ratio of dimeric populations, except for F466A, suggesting the relationship between the substrate transport activity and the formation of two distinct dimeric states (Fig. 5h). The F466A mutation did not inhibit dimer formation as much as other mutations, but its transport activity was quite low (Fig. 5g–h), indicating a crucial role of Phe466 in substrate transport. This mutation may affect the dimer interface rearrangement, rather than the dimer formation. The proper membrane expression of these hSVCT1 mutants was confirmed by observing fused GFP fluorescence using confocal microscopy (Supplementary Fig. 12), suggesting that destabilization of dimer formation do not affect trafficking of hSVCT1 to the plasma membrane, similarly to the fungal homolog UapA41. As a number of studies have provided evidence for the relationship between dimer formation and protein function of NAT family members20,21,23,42, these results further suggest the necessity of two distinct functional dimers to rearrange the interface into differently organized conformations through the transport cycle.
Structural comparison and conformational changes of dimers
To gain insights into the conformational changes of the dimer during the transport cycle, we compared our structures of hSVCT1 in two states with the previously reported dimeric UraA structure20, which is regarded as a representative of the substrate-bound occluded state of NAT/SLC23 family transporters, in accordance with the order of the predicted substrate transport cycle (Fig. 6a, Supplementary Fig. 13). When comparing the UraA structure in the substrate-bound occluded state with the hSVCT1 structure in the inward-open state, superimposed relative to the dimeric gate domains, the gate domains showed no significant movement between the two states, and thus probably maintain a stable dimer interface in this transition (Supplementary Fig. 13a). In contrast, the core domains exhibited a significant translocation toward the cytosolic side against the gate domains, with opening the substrate transport pathway between the core and gate domains inside each protomer (Supplementary Fig. 13b and Supplementary Movie 1). The bound substrate also shifts towards the cytosolic side through this transition (Supplementary Fig. 13a). This movement of the core domains is consistent with the typical elevator mechanism, as suggested in related structural reports21,24.

a Comparison of dimer structures of hSVCT1 and UraA between three states, in the order of putative substrate transport. Colored arrows indicate the relative movements of domains. b Schematic model for the predicted transport cycle of NAT/SLC23 members. Experimental structures are determined for the three colored states, and no structural information is currently available for the uncolored outward-open state.
In stark contrast, a comparison of our two hSVCT1 structures in the inward-open and substrate-free occluded states, superimposed relative to the core domains, revealed the distinct movement of the gate domain. In addition to the tilting movement observed in the protomeric comparison (Fig. 4f), the gate domain exhibited a horizontal shift relative to the core domain of the same protomer, possibly due to the structural rearrangement of the dimer interface discussed in the previous section (Supplementary Fig. 13c). As a result of this complex movement, the gate domain undergoes a rotational shift around the core domain of the same protomer and consequently approaches the core domain to occlude the transport pathway, while widely opening the cytosolic side of the interface between the dimers (Fig. 6a and Supplementary Movie 1). Simultaneously, the core domain slightly moves upward while maintaining a relatively rigid-body structure, in addition to the large shift of TM10 described in the protomeric comparison (Supplementary Fig. 13d). This is likely an extension of the elevator movement, which is augmented by the extensive movement of the adjacent gate domain and probably contributes to the transition to the next conformational state.
Interestingly, we found some lipid-like densities in both maps of the inward-open and occluded structures. We modeled phospholipid and cholesterol molecules, as well as the LMNG and CHS of detergent molecules, based on the shapes and sizes of the densities (Supplementary Figs. 6 and 14). Lipid-protein interactions have been suggested to contribute to the functional modulation or structural stabilization of transporter oligomers43,44. For instance, the lipid binding by the SVCT1-related proteins, UapA and prestin, was suggested to be important for dimerization and protein function23,29. The lipids are mainly localized on the protein surface over the boundary of the protomers in the inward-open structure (Supplementary Fig. 6). By contrast, in the substrate-free occluded structure, the widely opened inter-protomer cavity is occupied with multiple LMNG and CHS molecules, which seem to compress the gate domains (Supplementary Fig. 14). As the existence of lipid molecules at the dimer interface is consistent with the prediction for UapA in which lipids play a key role in the formation of functional dimers23, these LMNG and CHS molecules possibly mimic endogenous lipids and cholesterols, respectively. This notion is also supported by the fact that dimers are more stable under conditions containing LMNG than those containing DDM (Supplementary Fig. 1a, d), which can be ascribed to the difficulty of mimicking a phospholipid with two acyl tails by DDM with its single acyl chain.